Python Sorting Libraries and Algorithms Benchmark/Comparison

Sorting data is a basic task for data scientists and data engineers. Python users have a number of libraries to choose from with built-in, optimized sorting options. Some even work in parallel on GPUs. Surprisingly some sort methods don’t use the stated algorithm types and others don’t perform as expected.
Choosing which library and type of sorting algorithm to use can be tricky. Implementations change quickly. As of this writing, the Pandas documentation isn’t even up to date with the code (although my PR updating the sort options was just accepted).
In this article I’ll give you the lay of the land, provide tips to help you remember the methods, and share the results of a speed test.
Context
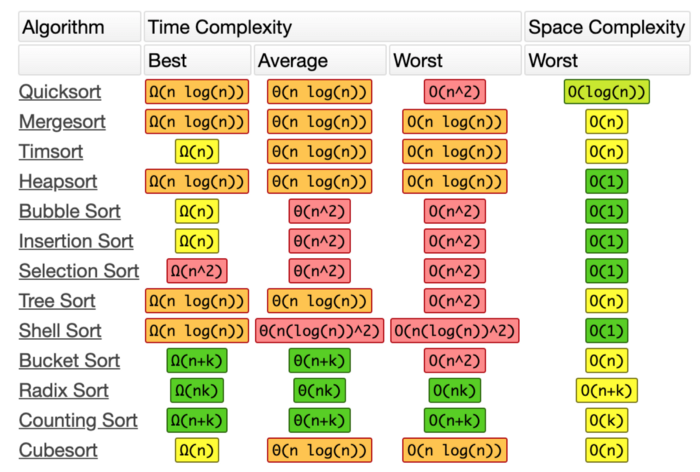
There are many different basic sorting algorithms. Some perform faster and use less memory than others. Some are better suited to big data and some work better if the data are arranged in certain ways. See the chart below for time and space complexity of many common algorithms.
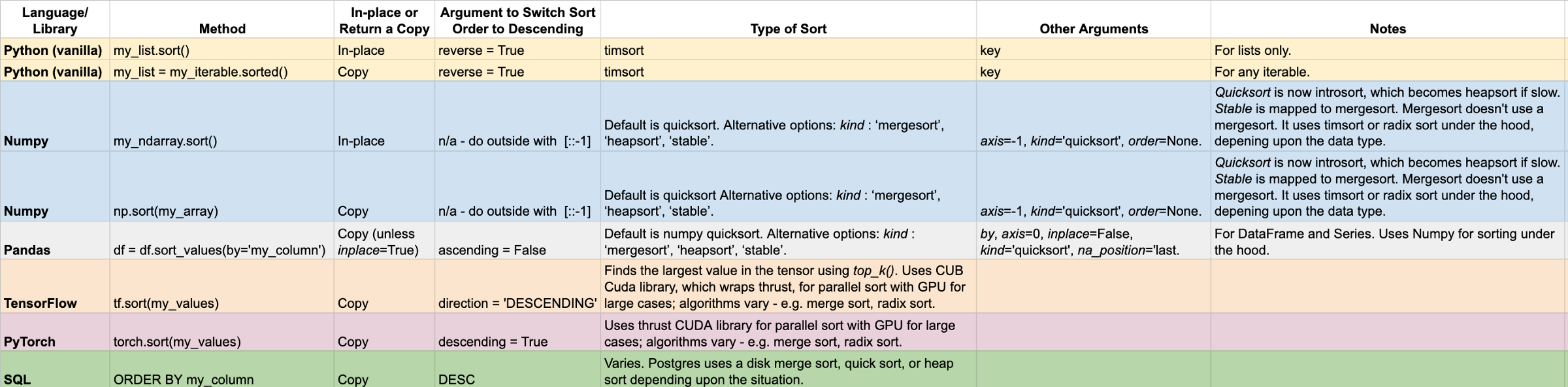
Being an expert at basic implementations isn’t necessary for most data science problems. In fact, premature optimization is occasionally sited as the root of all evil. However, knowing which library and which keyword arguments to use can be quite helpful when you need to repeatedly sort a lot of data. Here’s my cheat sheet.
The sorting algorithms have changed over the years in many libraries. These software versions were used in the analysis performed for this article.
python 3.6.8
numpy 1.16.4
pandas 0.24.2
tensorflow 2.0 beta
pytorch 1.1
Python (vanilla)
Python contains two built-in sorting methods.
-
my_list.sort()sorts a list in-place. It mutates the list.sort()returnsNone. -
sorted(my_list)makes a sorted copy of any iterable.sorted()returns the sorted iterable.sort()does not mutate the original iterable.
sort() should be faster because it is in place. Surprisingly, that’s not what I found in the test below. In-place sorting is more dangerous because it mutates the original data.
For vanilla Python all of the implementations we’ll look at in this article, the default sorting order is ascending — from smallest to largest. Most sorting methods accept a keyword parameter to switch the sort order to descending. Unfortunately for your brain, this parameter name is different for each library.
To change either sort order to descending in vanilla Python, pass reverse=True. key can be passed as a keyword argument to create your own sort criteria. For example, sort(key=len) will sort by the length of each list item.
The only sorting algorithm used in vanilla Python is Timsort . Timsort chooses a sorting method depending upon the characteristics of the data to be sorted. For example, if a short list is to be sorted, then an insertion sort is used. See Brandon Skerritt’s great article for more details on Timsort here .
Timsort , and thus Vanilla Python sorts, are stable . This means that if multiple values are the same, then those items remain in the original order after sorting.
To remember sort() vs. sorted(), I just remember that sorted is a longer word than sort and that sorted should take longer to run because it has to make a copy. Although the results below didn’t support the conventional wisdom, the mnemonic still works.
NumPy
Numpy is the bedrock Python library for scientific computing. Like vanilla Python, it has two sort implementations, one that mutates the array and one that copies it.
-
my_array.sort()mutates the array in place and returns the sorted array. -
np.sort(my_array)returns a copy of the sorted array, so it doesn’t mutate the original array.
Here are the optional arguments:
-
axis : int, optional — Axis along which to sort. Default is -1, which means sort along the last axis.
-
kind : {‘quicksort’, ‘mergesort’, ‘heapsort’, ‘stable’}, optional — Sorting algorithm. Default is ‘quicksort’. More on this below.
-
order : str or list of str, optional — When a is an array with fields defined, this argument specifies which fields to compare first, second, etc. A single field can be specified as a string, and not all fields need be specified, but unspecified fields will still be used, in the order in which they come up in the dtype, to break ties.
The sorting algorithms used are now a bit different than you might expect based on their names. Passing kind=quicksort means sorting actually starts with an introsort
algorithm. The docs
explain.
When [it] does not make enough progress it switches to a heapsort algorithm This implementation makes quicksort O(n*log(n)) in the worst case.
stable automatically chooses the best stable sorting algorithm for the data type being sorted. It, along with mergesort is currently mapped to timsort or radix sort depending on the data type. API forward compatibility currently limits the ability to select the implementation and it is hardwired for the different data types.
Timsort is added for better performance on already or nearly sorted data. On random data timsort is almost identical to mergesort. It is now used for stable sort while quicksort is still the default sort if none is chosen…‘mergesort’ and ‘stable’ are mapped to radix sort for integer data types.
From the Numpy docs
One take-away is that Numpy provides a wider range of control for sorting algorithm options than vanilla Python. A second take-away is that the kind keyword value doesn’t necessarily correspond to the actual sort type used. A final take-away is that the mergesort
and stable values are stable sorts, but quicksort
and heapsort
are not.
Numpy sorts are the only implementations on our list without a keyword argument to reverse the sort order. Luckily, it’s quick to reverse an array with a slice like this: my_arr[::-1].
The Numpy algorithm options are also available in the more user-friendly Pandas — and I find the functions easier to keep straight.
Pandas
Sort a Pandas DataFrame with df.sort_values(by=my_column). There are a number of keyword arguments available.
- by: str or list of str, required — Name or list of names to sort by. If axis is 0 or index then by may contain index levels and/or column labels. If axis is 1 or columns then by may contain column levels and/or index labels
- axis: {0 or index, 1 or columns}, default 0 — Axis to be sorted.
- ascending: bool or list of bool, default True — Sort ascending vs. descending. Specify list for multiple sort orders. If this is a list of bools, must match the length of the by argument.
- inplace: bool, default False — if True, perform operation in-place.
- kind: {quicksort, mergesort, heapsort, or stable}, default quicksort
— Choice of sorting algorithm. See also
ndarray.np.sortfor more information . For DataFrames, this option is only applied when sorting on a single column or label. - na_position: {'first', 'last'}, default 'last' — first puts NaNs at the beginning, last puts NaNs at the end.
Sort a Pandas Series by following the same syntax. With a Series you don’t provide a by keyword, because you don’t have multiple columns.
Because Pandas uses Numpy under the hood, you have the same nicely optimized sorting options at your fingertips. However, Pandas requires some extra time for its conveniences.
The default when sorting by a single column is to use Numpy’s quicksort . You’ll recall quicksort is now actually an introsort that becomes a heapsort if the sorting progress is slow. Pandas ensures that sorting by multiple columns uses Numpy’s mergesort . Mergesort in Numpy actually uses Timsort or Radix sort algorithms. These are stable sorting algorithms and stable sorting is necessary when sorting by multiple columns.
The key things to try to remember for Pandas:
- The function name:
sort_values(). - You need
by=column_nameor a list of column names. ascendingis the keyword for reversing.- Use mergesort if you want a stable sort.
When doing exploratory data analysis, I often find myself summing and sorting values in a Pandas DataFrame with Series.value_counts(). Here’s a code snippet to sum and sort the most frequent values for each column.
for c in df.columns:
print(f"---- {c} ---")
print(df[c].value_counts().head())
Dask , which is basically Pandas for big data, doesn’t yet have a parallel sorting implementation as of mid 2019, although it’s being discussed
Sorting in Pandas is a nice option for exploratory data analysis on smaller datasets. When you have a lot of data and want parallelized search on a GPU, you may want to use TensorFlow or PyTorch.
TensorFlow
TensorFlow is the most popular deep learning framework. The following is for TensorFlow 2.0:
tf.sort(my_tensor) returns a sorted copy of a tensor. Optional arguments:
- axis: {int, optional} The axis along which to sort. The default is -1, which sorts the last axis.
- direction: {ascending or descending} — direction in which to sort the values.
- name: {str, optional} — name for the operation.
tf.sort uses the top_k() method behind the scenes. top_k uses CUB
library for CUDA GPUs to make parallelism easier to implement. As the docs explain "CUB provides state-of-the-art, reusable software components for every layer of the CUDA programming model." TensorFlow uses radix sort on GPU via CUB, as discussed here
.
tf.sort() is a pretty intuitive method to remember and use if you work in TensorFlow. Just remember direction=descending to switch the sort order.
PyTorch
torch.sort(my_tensor) returns a sorted copy of a tensor. Optional arguments:
- dim: {int, optional} — the dimension to sort along
- descending: {bool, optional} — controls the sorting order (ascending or descending).
- out: {tuple, optional} — the output tuple of (Tensor, LongTensor) that can be optionally given to be used as output buffers.
Some digging showed that PyTorch uses a segmented parallel sort via Thrust if a dataset any larger than 1 million rows by 100,000 columns is being sorted.
Unfortunately, I ran out of memory when trying to to create 1.1M x 100K random data points via Numpy in Google Colab. I then tried GCP with 416 MB of RAM and still ran out of memory.
Segmented sort and locality sort are high-performance variants of mergesort that operate on non-uniform random data. Segmented sort allows us to sort many variable-length arrays in parallel. — https://moderngpu.github.io/segsort.html
Thrust is a parallel algorithms library that enables performance portability between GPUs and multicore CPUs. It provides a sort primitive that selects the most efficient implementation automatically. The CUB library used by TensorFlow wraps thrust. PyTorch and TensorFlow are using similar implementations for GPU sorting under the hood — whatever thrust chooses for the situation.
Like TensorFlow, the sorting method in PyTorch is fairly straightforward to remember: torch.sort(). The only tricky thing to remember is the direction of the sorted values: TensorFlow uses direction while PyTorch uses descending.
While sorting with GPUs could be a good option for really large datasets, it might also make sense to sort data directly in SQL.
SQL
Sorting in SQL is often very fast, particularly when the sort is in-memory.
SQL is a specification, but doesn’t dictate things like which sort algorithm an implementation must use. Postgres uses
a disk merge sort, heap sort, or quick sort, depending upon the circumstances. If you have enough memory, sorts can be made much faster by making them in-memory. Increase the available memory for sorts via the work_mem setting.
Other SQL implementations use different sorting algorithms. For example, Google BigQuery uses introsort with some tricks, according to this Stack Overflow answer .
Sorts in SQL are performed with the ORDER BY command. This syntax is distinct from the Python implementations that all use some form of the word sort. I find it easier to remember that ORDER BY goes with SQL syntax because it’s so unique.
To make the sort descending, use the keyword DESC. So a query to return customers in alphabetical order from last to first would look like this:
SELECT Names FROM Customers ORDER BY Names DESC;
Comparisons
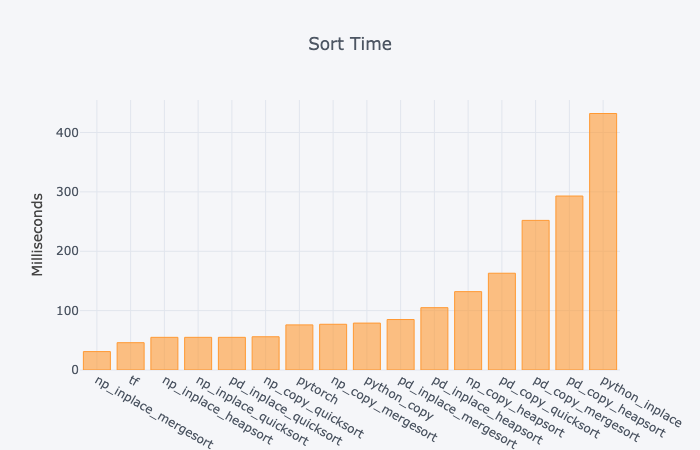
For each of the Python libraries above, I conducted an analysis of the wall time to sort the same 1,000,000 data points in a single column, array, or list. I used a Google Colab Jupyter Notebook with a T4 GPU.
Observations
- For both Numpy and Pandas, inplace is faster than copying the data. No surprise there.
- The default Pandas quicksort is rather fast.
- Most Pandas functions are comparatively slow.
- TensorFlow is quite fast.
- Python inplace sorting is surprisingly slow. It was 10x slower than the Numpy inplace mergesort or the TensorFlow sort. I tested it multiple times (with different data) to double check that this was not an anomaly.
Again, this is just one small test. It’s definitely not definitive.
Wrap
You generally shouldn’t need custom sorting implementations. The off-the shelf options are strong. They are generally not using just a single sorting method. Instead they evaluate at the data first and then use a sorting algorithm that performs well. Some implementations even change algorithms if the sort is not progressing quickly.
In this article, you’ve seen how to sort in each part of the Python data science stack and in SQL. I hope you’ve found it helpful. If you have, please share it on your favorite social media so others can find it, too.
You just need to remember which option to choose and how to call them. Use my cheat sheet above to save time. My general recommendations are the following:
- Use the default Pandas
sort_values()for exploration on relatively small datasets. - For large datasets or when speed is at a premium, try Numpy’s in-place mergesort, a PyTorch or TensorFlow parallel GPU implementation, or SQL.
Sorting on GPUs isn’t something I’ve seen much written about. It’s an area that appears ripe for more research and tutorials. Here’s a 2017 article to give you a taste of recent research . More info on GPU sorting algorithms can be found here .